Dietary Interactions in Ecology Through Sequencing (DIETS) Symposium
21st – 24th July 2026

Trophic interactions underpin the function, stability and diversity of our ecosystems, rendering their study vital for understanding, protecting and monitoring biodiversity. The last two decades have seen the rise and dominance of molecular methods for the analysis of wild animal diets, with widening adoption introducing greater diversity in practices, and opportunities for advancement and collaboration.
The Dietary Interactions in Ecology Through Sequencing (DIETS) Symposium, to be held at Durham University 21st-24th July 2026, will be an in-person symposium focused on the molecular analysis of trophic interactions. By sharing recent and ongoing research and ideas, DIETS will catalyse collaboration and advancement in this field whilst also identifying consensus on various practical aspects of molecular dietary analysis.


DIETS will take place across four days, including a one-day workshop, a local excursion and two days of talks from global researchers using molecular methods to study trophic interactions. Register or submit an abstract using the buttons below!
General registration deadline: 17:00 BST, 26th June 2026.
Registration fee: £195 standard rate and £160 student rate.
Workshop fee: £35, capped at 30 attendees (first come, first served).
Excursion fee: £35, capped at 40 attendees (first come, first served).
Abstract deadline: 17:00 BST, 1st May 2026.
We are keen to receive submissions from speakers across a broad range of disciplines, career stages and personal backgrounds, especially those from underrepresented or historically excluded groups. We will evaluate and improve the inclusivity of our programme throughout and welcome any feedback.
If you have any questions about the event, please feel free to reach out to Jordan Cuff or Andreanna Welch.
Plenary speakers

Michael Traugott
Universität Innsbruck, Austria
“25 years of analysing trophic interactions molecularly: key learnings and future prospects”

Vanessa Mata
Cibio, Portugal
“DNA metabarcoding of small vertebrate diets: promises, pitfalls and ways forward”

Simon Creer
Prifysgol Bangor, UK
“What can environmental DNA tell us about biodiversity and ecological interactions across diverse biomes?”


Event sponsors


Schedule
21st July: Workshop on containerised bioinformatics for dietary metabarcoding via nanopore sequencing
Hosted by Dr Chris Wyatt and Fernando Duarte Frutos (Eco-Flow Team, University College London)
The advancement of molecular dietary analysis has closely tracked developments in sequencing technology. Nanopore sequencing represents one of the most widely acclaimed advances of recent years, with its associated long-read sequencing being awarded Nature‘s ‘method of the year 2022’. The variable read length capabilities of nanopore sequencing, paired with its low capital costs and portability, make it an accessible and attractive prospect, yet it remains scarcely applied to molecular dietary analysis, in part due to its divergent bioinformatic needs.
The Eco-Flow team at University College London, led by Dr Chris Wyatt, have developed a containerised (i.e., semi-automated) bioinformatics pipeline for the analysis of nanopore metabarcoding data. Eco-Flow is a BBSRC-funded project aiming to make bioinformatics more accessible and usable for agri-ecology and the wider scientific community.
This workshop will introduce, demonstrate and provide hands-on experience with running this new nanopore metabarcoding bioinformatics pipeline. The workshop will provide an introduction to running nf-core pipelines locally and on high-performance computing clusters (HPCs), alongside debugging pipelines and the basics of nextflow. All of this knowledge will then be applied to running a practical example of nf-core based on both RNA-Seq data and the new metabarcoding pipeline.
Attendees are encouraged to bring along a laptop to run parts of the workshop themselves. Attendance at the workshop costs an additional £35 to cover catering and refreshments costs for the day. Places are limited to 25 on a first-come-first-served basis.

22nd July: Symposium
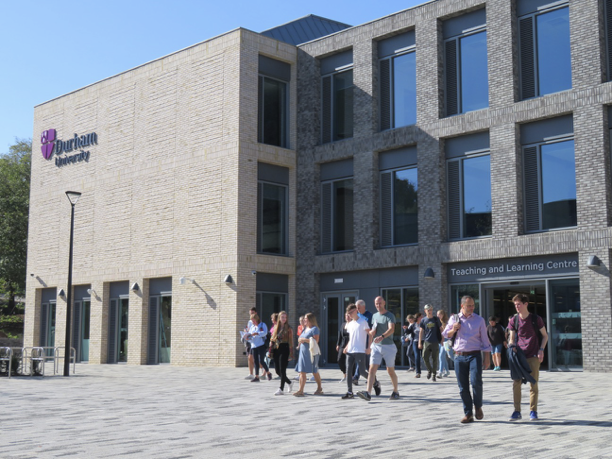
All sessions of the main symposium will take place in the Teaching and Learning Centre, Durham University, unless otherwise stated (in italics).
| Time | Activity |
| 9:00 | Registration |
| 9:30 | Welcome and housekeeping |
| 10:00 | Talk session |
| 11:00 | Break |
| 11:30 | Breakout session 1 |
| 12:30 | Lunch |
| 13:30 | Poster session |
| 14:00 | Plenary 1 |
| 14:30 | Talk session |
| 15:30 | Break |
| 16:00 | Breakout session 2 |
| 17:00 | Mixer and poster session |
| 18:30 | Plenary 2, Hatfield College Chapel |
| 19:00 | Plenary panel discussion, Hatfield College Chapel |
| 19:30 | Conference dinner, Hatfield College Dining Hall |
23rd July: Symposium
All sessions of the main symposium will take place in the Teaching and Learning Centre, Durham University, unless otherwise stated (in italics).
| Time | Activity |
| 9:00 | Talk session |
| 10:30 | Break |
| 11:00 | Breakout session 3 |
| 12:00 | Lunch |
| 13:00 | Poster session |
| 14:00 | Plenary 3 |
| 14:30 | Talk session |
| 16:00 | Break |
| 16:30 | Breakout summaries and open discussion |
| 17:00 | Conference close |
Breakout sessions
For the breakout sessions, three parallel facilitated discussions will take place. Attendees will have the opportunity to submit their preferences for each breakout session. The goal is to discuss current challenges, opportunities and best practices related to each of these themes. No prior knowledge is required, all experience levels are encouraged to contribute to discussions and you do not need to have worked on a specific topic to contribute.
| Groupings | Breakout session 1 | Breakout session 2 | Breakout session 3 |
| Group 1 | Predatory dietary analysis | Experimental controls | Reference data |
| Group 2 | Herbivorous dietary analysis | Data processing and filtering | Open research and FAIR |
| Group 3 | Omnivorous dietary analysis | Merging methods | Multi-marker metabarcoding |
Example topics to be covered in each breakout session:
- Predatory dietary analysis: The analysis of predator diets using molecular methods offers powerful insights into dynamic animal-animal interactions. There are many challenges associated with this approach though, especially due to the phylogenetic proximity of predator and prey in many cases. This breakout session will cover these challenges and ways of overcoming them (relevant example paper: Cuff et al., 2023)
- Herbivorous dietary analysis: Detecting plants (and other sessile resources) in the diets of consumers is a widespread application of molecular dietary analysis. It is not, however, without its specific problems, from variable DNA densities in plant resources to highly prevalent surface contaminants, which this breakout session will cover.
- Omnivorous dietary analysis: The analysis of omnivorous diets using techniques like DNA metabarcoding is crucial for understanding the various direct and indirect interactions between plants, animals and beyond. It is not, however, without its specific challenges, particularly linked to secondary consumption and false inferences, which this breakout session will cover (relevant example paper: Tercel et al., 2021).
- Experimental controls: Stringent molecular dietary analysis workflows often include experimental controls, such as negative controls, blanks, positive controls, mock communities or spike-ins. These can help identify contaminants, biases and other shortfalls when using methods like DNA metabarcoding, but there are inconsistencies in their application. This breakout session will cover different types of controls, what they show and how they are currently used (relevant example paper: Zinger et al., 2019).
- Data processing and filtering: Molecular methods, due to their incredible sensitivity, are often susceptible to contamination and other false positive detections. Addressing these detections can be complex and often results in false negatives (i.e., missing data). This breakout session will cover this balancing act and the promises and pitfalls involved in filtering molecular dietary data (relevant example papers: Littleford-Colquhoun et al., 2022; Drake et al., 2022).
- Merging methods: Sometimes molecular dietary analysis alone isn’t enough to collect the data we need, or using it in combination with other methods can vastly improve our insights into the natural world. Merging different data types together is not, however, straightforward, and there are many challenges and considerations involved, which this breakout session will cover (relevant example papers: Evans et al., 2016; Cuff et al., 2022).
- Reference data: In order to relate sequences to taxonomic identities, we rely on reference databases of sequences from identified samples. These reference data can, however, be mislabelled, poorly resolved taxonomically, or contain sequencing errors or sequence/taxonomic conflicts. Many taxa are entirely missing from databases, which often underrepresent intraspecific variation, too. This breakout session will cover these challenges and the pathways to overcoming them (relevant example paper: Keck et al., 2023).
- Open research and FAIR: As we generate more and more sequencing data, it’s crucial that this is made accessible to all. Making our data and the processes underpinning it findable, accessible, interoperable and reproducible is key to ensuring we use, analyse and re-use data responsibly, but there are challenges involved in making this streamlined, effective and future-proof, which this breakout session will cover (relevant example paper: Takahashi et al., 2025).
- Multi-marker metabarcoding: The parallel use of multiple PCR primer pairs is a valuable approach to mitigating PCR biases and targeting multiple distinct taxonomic groups. It is not, however, without its complications and costs. This breakout session will discuss these nuances and best practices (relevant example paper: da Silva et al., 2019).
24th July: Excursion to WWT Washington
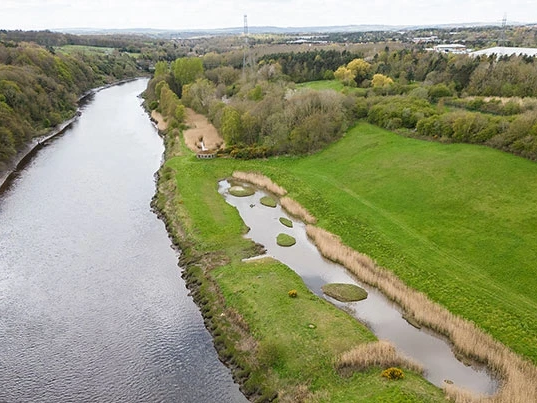
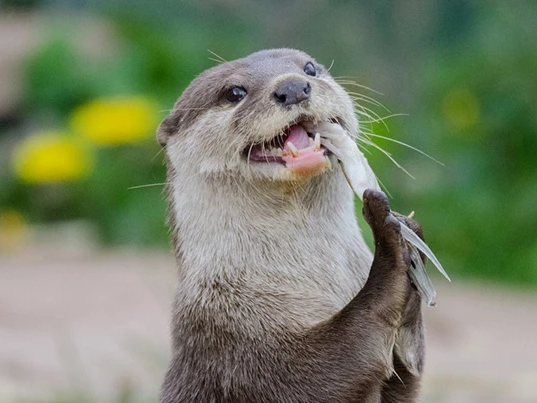
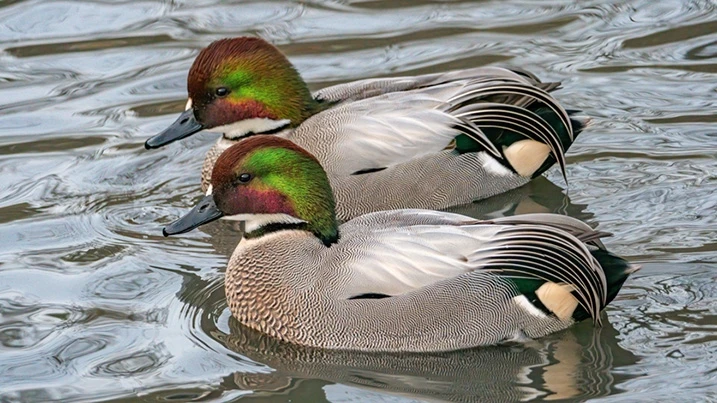
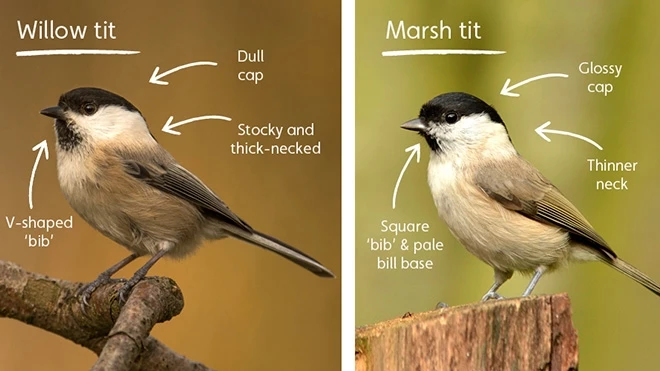
An optional excursion to Washington’s Wetland Centre, a premier wetland conservation centre, will be run on the final day. This will be an ideal time to reflect on the conference in the beautiful surroundings of a thriving wetland landscape. Watch trophic interactions unfold before your eyes (rather than in a sequencer) between a range of birds, arthropods and small mammals. Chilean flamingos, otters, willow tits and a wide range of waterbirds can be found throughout the site, representing wildlife from across Britain and beyond.
There will be opportunities to watch otter feeding and other guided activities for close-up encounters with some fantastic wildlife, including pond dipping for aquatic invertebrates. The site boasts a range of interesting habitats, including a saline lagoon on the banks of the river Wear. Visiting in July, the peak for wetland biodiversity and activity, will provide plenty of opportunity to relate the topics of the conference to the natural world, whilst also deepening your connections with other delegates.
Attendance for the excursion costs an additional £45 to cover catering and travel costs from Durham. Places are limited to 35 on a first-come-first-served basis. Additional attendees can join on the day if they can arrange transport and catering and cover their entrance fee privately.




Organising Committee